This article gives you a guide on how to deploy Spark RAPIDS to GCP Dataproc clusters and optimize it.
NVIDIA has developed a Spark plugin for GPU Acceleration called the RAPIDS Accelerator for Apache Spark. In a previous article, we showed you how Spark RAPIDS works, how to apply it to your scripts, and how to set it up on Databricks.
In this article, we will show you how to deploy Spark RAPIDS to GCP Dataproc clusters, how to properly optimize your configuration for maximum performance, and share the results of benchmarks we performed that compared GPU accelerated against regular clusters.
Overview
Over the last decade, the amount of data computed in software projects exploded. This ever-increasing amount of data, coupled with issues of data quality and variety, lead to the rise of production-grade Big Data frameworks, like Apache Spark, to process these large amounts of data effectively. Today, such data pipelines are often hosted in the cloud where consumption-based cost models apply. Therefore, optimizing the runtime performance of these software applications becomes critical for reducing their total cost of ownership.
In the area of deep learning, GPU acceleration has led to significant performance improvements for model training. However, a production read data product does not only consists of model training and inference but also of many data processing and transformation steps that could also benefit from gpu acceleration. For this purpose, NVIDIA has developed a series of open source libraries that apply GPU acceleration to other data science tasks like data preparation and ETL. These libraries follow the APIs of well-known data science libraries like Pandas, but an underlying CUDA implementation delivers native GPU performance, similar to how GPU acceleration for TensorFlow works. The RAPIDS Accelerator for Apache Spark makes use of these libraries to define new optimization rules for the Spark Catalyst Query Optimizer. This approach allows Spark Jobs to be run on GPUs without any code changes. Given the performance improvements, GPU acceleration has provided for deep learning, correctly applying it to Spark Jobs could also lead to significant speedups and cloud cost savings.
In this blog post, we will cover:
- How to set up Spark RAPIDS on GCP Dataproc
- How to properly configure Spark RAPIDS to obtain optimal results
- Concrete speedup results measured on the TPCx-BB benchmark as well as own, specialized transcoding and repartitioning benchmarks
Prerequisites for following along are a basic understanding of Apache Spark as well as institutional access to GCP with the permission to provision VMs with T4 GPUs.
Setting up Spark RAPIDS on Dataproc
The first step necessary to run Spark RAPIDS on Dataproc is provisioning a GPU cluster and installing the RAPIDS libraries on that cluster. Open up the Google Cloud console and navigate to a project of your choice, or set up a new project to test Spark RAPIDS.
Search for “Dataproc“ in the top bar. This will lead you to the cluster overview. Click on the “Create Cluster“ button to begin the setup process.

When prompted in a popup to select the cluster type, choose “Cluster on Compute Engine“. Select your preferred region and give the cluster a name. Use a standard cluster and disable autoscaling. Activate “Component Gateway“ to access the Spark UI.
Next, configure the nodes of the cluster. For the master node, a simple 8 or 16-core standard node is sufficient. Use 16 core nodes of the highmem type for your worker nodes with additional RAM. Select “CPU Platform and GPU“ and add one T4 GPU per node. The number of nodes you should use is dependent on the workload size. In our experiments, a cluster with four worker nodes was able to compute workloads up to 1 terabyte of input in a reasonable time.

Under “Customize cluster“, make sure to switch the YARN capacity scheduler to the resource scheduler by adding a property with “capacity-scheduler“ as the prefix, “yarn.scheduler.capacity.resource-calculator“ as the key, and “org.apache.hadoop.yarn.util.resource.DominantResourceCalculator“ as the value.

Otherwise, the default scheduler will override the Spark executor configuration we will specify later to maximize performance. Under initialization actions, register the two setup scripts offered by Google here and here by copying them to a Cloud Storage Bucket and selecting them in the dialogue. These scripts set up CUDA and RAPIDS on the cluster. Your cluster is now ready to deploy and should be up and running in five to seven minutes.
To execute a Spark job on the cluster, open the “Jobs“ tab. Click on “Send job“ in the toolbar. Name your job and select the cluster you just created to execute the job on. Upload the JAR or Python file to a Cloud Storage Bucket, and select the appropriate job type and the file (for example, you could use one of the sample jobs for Spark RAPIDS provided by NVIDIA). The Spark configuration settings can be added in the “Attributes“ section. We will now have a look at those settings and show you how to optimize them.
Configuring a Spark RAPIDS job
The necessary configuration parameters can be split into two categories: general Spark settings and Spark RAPIDS-specific settings.
spark.Executor.instances determines how many executor processes will be created with the cluster’s resources. For the use of RAPIDS, the number of executors must be equal to the number of GPUs. In our example cluster, this would lead to four executors.
spark.Executor.cores controls the number of CPUs per executor. Since the number of Executors is fixed to the number of GPUs in our case, it is best to equally distribute the available CPUs over the executors. In our example case, this would mean using 15 cores per executor and reserving one CPU per node for the OS.
spark.Executor.memory describes how much RAM is assigned to the executors. As with the CPU cores, distribute the memory equally among the executors for the best results, while reserving around 4 GB per node as overhead.
spark.sql.shuffle.partitions determines how many partitions will be used to transfer data between nodes during shuffle steps. It is best to leave this number up to Spark’s Adaptive Query Execution mechanism for optimal results. Use the default value of shuffle partitions and make sure spark.sql.adaptive.coalescePartitions.enabled is set to true.
spark.sql.files.maxPartitionBytes controls the maximum partition size obtained when reading a file. When using GPUs, we generally want larger partitions to make the most of the GPU’s high throughput. For T4 GPUs, 1 GB is a good partition size.
spark.Executor.resource.gpu.amount sets the number of GPUs per Executor. For Spark RAPIDS, set this to one GPU per executor, since the Framework only supports a 1:1 mapping of Executors and GPUs.
spark.task.resource.gpu.amount controls how many tasks can be run simultaneously on a cluster with a GPU. It does not control how many tasks run on the GPU itself. Set this to one divided by the number of cores per executor, resulting in 1/15 on our example cluster.
spark.rapids.sql.concurrentGpuTasks determines how many tasks can be executed in parallel on the GPU. If this number is lower than spark.task.resource.gpu.amount, tasks will compute their CPU operations and then enter a queue as soon as they require the GPU. The ideal number of parallel tasks depends on the GPU type. When using T4 GPUs, two tasks per GPU obtain optimal results.
spark.rapids.memory.pinnedPool.size controls how much memory will be pinned, preventing the OS from relocating it or swapping it to disk. This memory can then be used by the GPU to transfer data between its memory and the main memory asynchronously from the CPU, greatly increasing performance. In our example configuration, set this value to the 4 GB we reserved during executor memory allocation.
spark.rapids.sql.batchSizeBytes determines the maximum size of the columnar batches the GPU computes. Match this value to spark.sql.files.maxPartitionBytes, which leads to 1 GB for our example cluster.
spark.rapids.memory.gpu.pool controls which memory pooling algorithm will be used by RAPIDS. Memory pooling allows RAPIDS to view the memory of the different GPUs on the cluster as a single resource. Set this value to “ARENA“ to use the ARENA algorithm. This algorithm prioritizes reducing fragmentation over allocation performance, which is more beneficial for the GPU since the memory can already be accessed much faster than RAM.
Aside from these settings which mostly concern themselves with hardware allocation, there is a series of settings that prevent certain operations to run on the GPU by default because the results might not match the results obtained when executing them on the CPU. Since the differences are minuscule, they can be safely ignored in most use cases. It is, therefore, crucial to enable these operations to be run on the GPU for maximum performance to eliminate CPU fallbacks. CPU fallbacks occur when an operation cannot be computed on the GPU, either because it is not yet implemented or the configuration prohibits it from being run on the GPU. These fallbacks are extremely detrimental to performance since they require writing data from the GPU’s memory to the main memory, using the slower CPU to compute them, and then writing the result back to the GPU’s memory to continue computation. This introduces a massive I/O overhead that would not exist if the operation would have been performed on the GPU. It is therefore vital to set the following configuration values to true:
- spark.rapids.sql.incompatibleOps.enabled
- spark.rapids.sql.variableFloatAgg.enabled
- spark.rapids.sql.castFloatToDecimal.enabled
- spark.rapids.sql.castFloatToIntegralTypes.enabled
- spark.rapids.sql.exec.CollectLimitExec
- spark.rapids.sql.castStringToFloat.enabled
The following table provides a full overview of all the important settings and their recommended values, assuming the use of T4 GPUs.
| Setting | Value |
|---|---|
| spark.Executor.instances | Number of GPUs in the cluster |
| spark.Executor.cores | Distributed equally over the executors, reserving 1 per node for OS |
| spark.Executor.memory | Distributed equally over the executors, but reserve 4GB per node |
| spark.sql.shuffle.partitions | 200 |
| spark.sql.adaptive.coalescePartitions.enabled | true |
| spark.sql.files.maxPartitionBytes | 1 GB |
| spark.Executor.resource.gpu.amount | 1 |
| spark.task.resource.gpu.amount | 1 / spark.Executor.cores |
| spark.rapids.sql.concurrentGpuTasks | 2 |
| spark.rapids.memory.pinnedPool.size | 4 GB |
| spark.rapids.sql.batchSizeBytes | 1 GB |
| spark.rapids.memory.gpu.pool | ARENA |
| spark.rapids.sql.incompatibleOps.enabled | true |
| spark.rapids.sql.variableFloatAgg.enabled | true |
| spark.rapids.sql.castFloatToDecimal.enabled | true |
| spark.rapids.sql.castFloatToIntegralTypes.enabled | true |
| spark.rapids.sql.exec.CollectLimitExec | true |
| spark.rapids.sql.castStringToFloat.enabled | true |
Benchmarking Spark RAPIDS
We ran several benchmarks to measure the performance increase offered by Spark RAPIDS on Dataproc clusters. Among these benchmarks was the industry standard TPCx-BB benchmark, which contains thirty queries executed on a mix of structured, semi-structured, and unstructured data, reflecting the variety faced by most data science projects. The TPCx-BB specification also includes machine learning and NLP operations, but Spark RAPIDS currently only accelerates the Spark SQL module. Therefore, we developed an adapted version of the TPCx-BB benchmark. This version only contains the purely analytical queries as well as the ETL and preparation portions of the machine learning and NLP operations which can be executed using Spark SQL. On a 1 TB scale, one query ran into OOM issues. While these issues can be addressed by modifying the configuration, the necessary changes would degrade the performance. Therefore we removed the query from the 1 TB workload and only ran the remaining 29 queries. We furthermore developed benchmarks that measure the performance of transcoding data from CSV to Parquet as well as the repartitioning of DataFrames. For all experiments described below, the GPU cluster consisted of four nodes of the n1-highmem-16 type with one T4 per node for acceleration, and the CPU cluster was made up of six 16-core high memory nodes. This resulted in an hourly cost of 5.28€ for the GPU accelerated cluster and 5.96€ for the CPU cluster in the europe-west-4 GCP region. The software configuration matched the example configuration outlined above.
Below you can see the mean execution time of running TPCx-BB on one terabyte of input data on both clusters. GPU acceleration halved the runtime in this case. The more complex an operation was, the greater the runtime reduction was, with high cardinality JOIN operations and complex WHERE clauses benefiting the most.
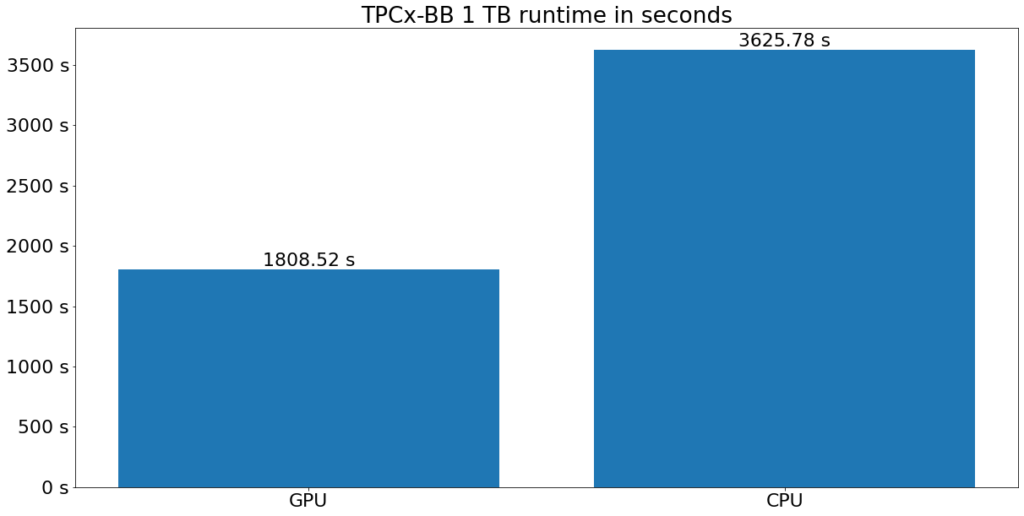
This runtime reduction also leads to reduced execution costs since Dataproc only charges for running resources. The chart below shows the execution cost of the benchmark on both clusters. The execution cost is more than halved with GPU acceleration.

We executed our TPCx-BB implementation with varying data sizes to verify whether Spark RAPIDS delivers increasing performance benefits with more data. Below you can see a chart that plots the execution times on CPU and GPU for 1 GB, 10 GB, 100 GB, and 1 TB. With rising amounts of data, the runtime advantage of the GPU cluster increased.

The transcoding benchmark measured the runtime of transcoding multiple CSV files into Parquet files. The chart below shows the runtimes on both clusters when transcoding one terabyte of data. GPU acceleration led to a 39 % runtime reduction. The runtime reductions for the individual CSV files correlated with their size: the bigger the file, the greater the runtime reduction.

The execution costs for the one terabyte transcoding experiment are shown below.
GPU acceleration obtained a 45 % cost reduction.

The repartitioning benchmark tested loading a set of Parquet files of varying schemas and sizes from a Cloud Storage Bucket, repartitioning them into 4, 20, and 100 target partitions before writing them back into the Cloud Storage Bucket. respectively. The chart below shows the runtimes on one terabyte of input data. With 4 target partitions, GPU acceleration reduces the runtime by 75 %. With 20 target partitions, the runtime is reduced by 57 %. Finally, with 100 target partitions, the runtime is reduced by 27 %. GPU acceleration leads to incredible performance advantages with a low number of input partitions, and still outperforms the CPU with a high number of target partitions. By analyzing the runtimes for the separate files, we again determined that larger files benefit more than smaller ones.

The execution costs for the repartitioning experiment are shown below. The execution cost on the GPU cluster is reduced by 77 % for 4 target partitions, 62 % for 20 target partitions, and 35 % for 100 target partitions.
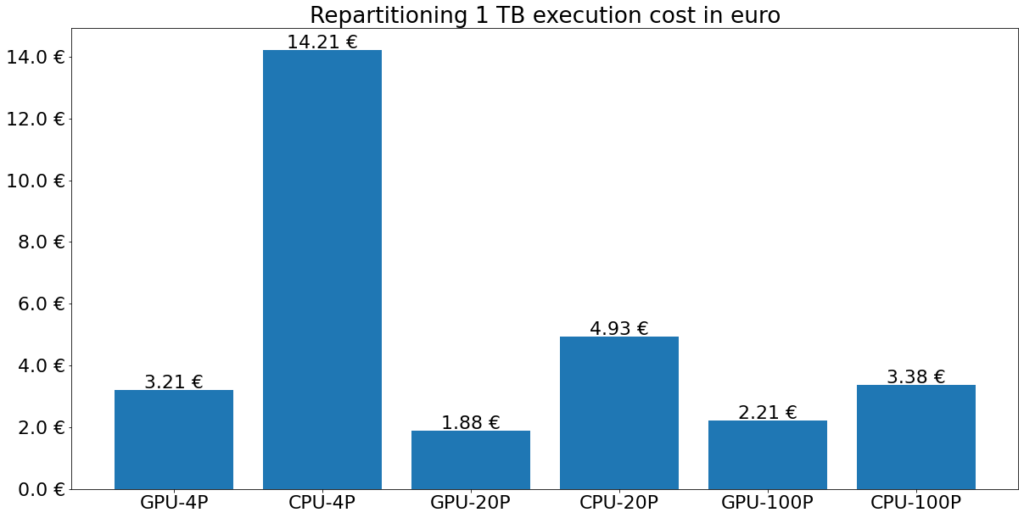
Conclusion
Across all our experiments, GPU acceleration resulted in a clear runtime advantage which also led to reduced execution costs, as long as the outlined configuration was used. No changes to the code had to be made to any of the scripts to run them on the GPU, making the adoption of Spark RAPIDS fairly straightforward.
As mentioned before, in certain situations, particularly high cardinality JOIN operations on large amounts of data, OOM errors can occur. These types of errors can be fixed by scaling down certain configuration parameters. The first thing to try is lowering spark.rapids.sql.concurrentGpuTasks if multiple tasks on the GPU are running in parallel. If the OOM errors still occur after lowering spark.rapids.sql.concurrentGpuTasks to 1, lower spark.rapids.sql.batchSizeBytes and spark.sql.files.maxPartitionBytes. This resolved the OOM issues in our case. However, lowering these parameters leads to performance degradation since throughput and parallelism are impacted by them. Even under these conditions, the GPU clusters produced better results than the CPU clusters, but the acceleration was not as high as it could have been. Also, even with all operations enabled, certain queries resulted in fallbacks to the CPU due to operators not yet being implemented, though this will likely be mitigated in the future as more and more operators are implemented.
In addition to acceleration with T4 GPUs, we also ran experiments using the larger A100 GPUs. While those GPUs lead to even faster computation, their cost-to-performance ratio is too low to lead to tangible improvements over a CPU cluster with similar hourly costs.
Overall, using Spark RAPIDS was straightforward since no code changes were necessary, and preexisting Spark jobs could simply be run with the configuration outlined above. Spark RAPIDS currently only accelerates Spark SQL, acceleration for MlLib, Streaming, and GraphX are pending. With an average runtime reduction of 34 % and a resulting average cost reduction of 42 %, Spark RAPIDS is a technology worth exploring.



