Generative AI has experienced astonishing progress over the course of the past years. Especially the improvements of multi-modal modeling lead to great excitement in the field. Within this blog post we will discuss the adaptation of the multi-modal model CLIP to the chest radiography domain using the MIMIC-CXR dataset.
The CLIP models (Blog, Paper) work well on data of the general domain, but require adaptation for more specialized tasks. The abundance of training data for the CLIP models, basically a large amount of publicly available text-image pairs on the internet, yielded great results across many general domain benchmarks, such as CIFAR or ImageNET. When it comes to more specialized tasks, like chest radiographs, features of the pretrained CLIP do not transfer well to this domain. Within this blog post we share some of our learnings of using contrastive language supervision to perform domain adaptation:
Contrastive Language Supervision successfully adapts CLIP to the domain of chest X-rays
Our assumption is that CLIPs pretrained features do not transfer well to the domain of chest X-rays and that additional adaptation to the respective domain would improve the models performance. For this reason, we compare domain adaptation using contrastive language supervision (CLS) with plain supervised fine-tuning (FT) on labels and use AUC as the metric to compare our approaches. We find that CLS combined with linear probing performs better than only using FT on labels.
The MIMIC-CXR dataset contains over 227,000 chest radiograph studies with accompanying expert reports. The labels for the studies were extracted with a label extractor from the CheXpert dataset, and 14 diagnoses were used, including pneumonia. The images were resized such that the smaller side had a length of 256 pixels.
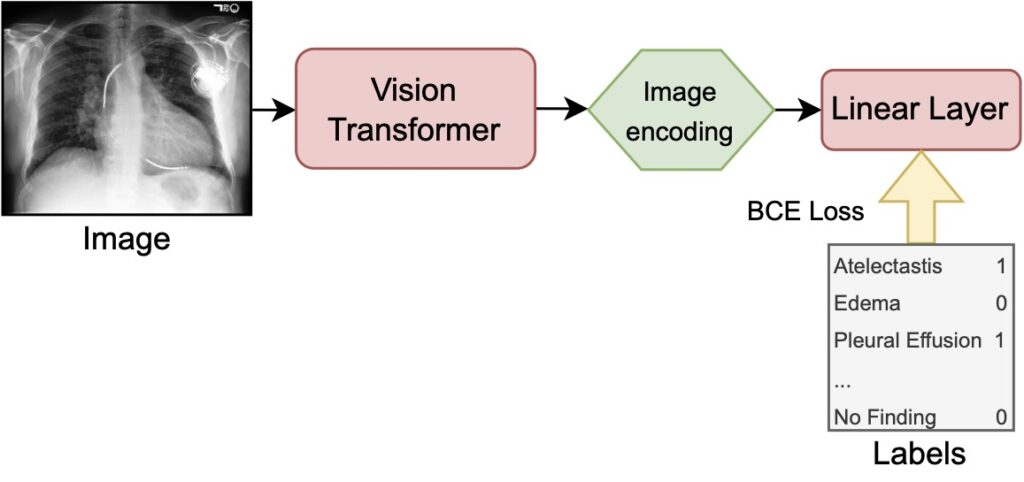
Fine-Tuning on labels (FT) leverages the pretrained vision transformer to encode images (our X-ray scans) to latent representations and passes them to a linear layer in order to perform a multi-label classification. During training the full model is optimized, meaning that the classification layer as well as the image encoder weights get adjusted. The binary cross entropy (BCE) loss is calculated per label and then averaged over all labels yielding the supervised loss used as the training objective.
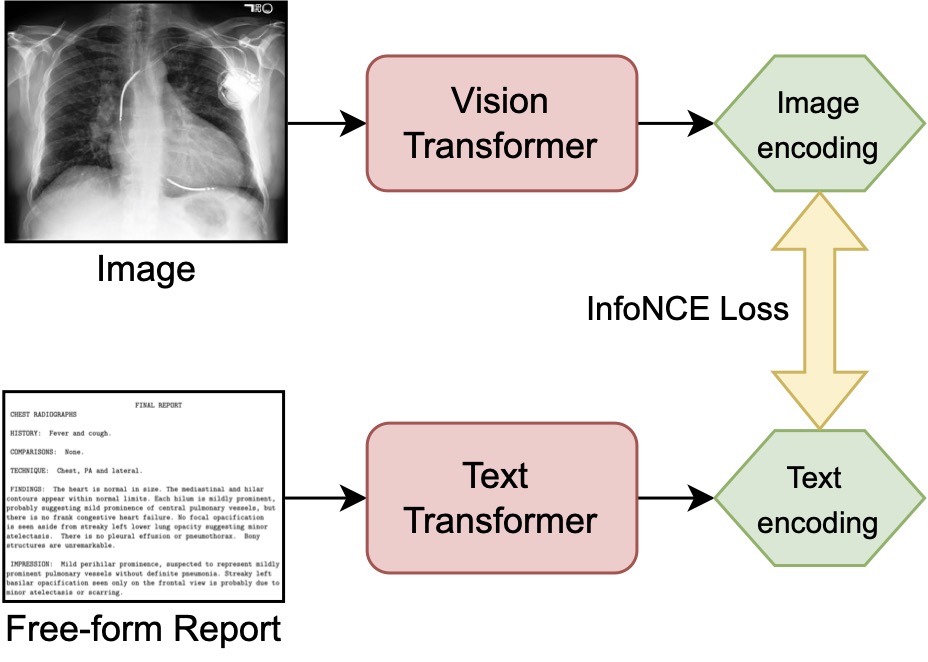
Contrastive Language Supervision (CLS) uses images (our X-ray scans) and texts (corresponding reports) as inputs to the pretrained vision and text transformers. Both networks encode their respective inputs to a latent representation. Both representations are aligned by training with the InfoNCE loss. After this adaptive pretraining step, a linear probe is applied to receive a prediction on the downstream task and compare it to the results of FT on the ground truth labels.
First, we find that features of the general CLIP model do not transfer well to the chest radiographs of MIMIC-CXR. Applying a linear probe on top of the pretrained weights yields an AUC of 66.7, which is only slightly better than randomly initialized weights combined with a linear probe (AUC: 66.5). Both approaches are significantly outperformed by models making use of domain specific data.
Second, we find that pretraining CLIP with CLS successfully adapts the model to the domain of chest X-rays. CLS combined with a linear probe (AUC 77.8) is competitive with and even slightly superior to CLS plus FT (77.3) and pure FT (AUC 77.2). The results of our dataset size ablation study show that pretraining using CLS on the whole dataset, followed by fine-tuning on a fraction of the labels consistently performs best.
Domain Adaptation with Contrastive Language Supervision generalizes to other datasets of the same domain and works well on limited data
The RSNA Pneumonia and cheXpert datasets were used to evaluate if the CLIP model pretrained on MIMIC-CXR generalizes to data from other hospitals with other labels. Linear probes were trained on the features of the pretrained models to predict the labels of the external datasets. The cheXpert dataset contains 223,648 images labeled with the same diagnoses as MIMIC-CXR. The official validation split was used as our test set, as there is no official test set released. The RSNA dataset contains 30,227 images of which 9,555 are annotated with the pneumonia diagnosis, forming a single-class, single-label classification task. A random subset of 10% of the data was used as a test set, the rest was used for training.
We find that our domain adapted CLIP is competitive to state-of-the-art approaches on both datasets and conclude that CLS generalizes well on datasets of the same domain. On cheXpert, our CLS-adapter model with linear probe scores an AUC of 87.2, which is on a similar level as results reported by Zhang et al. (2020). Interestingly, their best approaches combine CLS with linear probing (AUC 87.3) and CLS combined with FT (AUC 88.1) as well. Our model outperforms other reported approaches from Azizi et al. (2021), who use FT (AUC 77.0), and Seibold et al. (2022), who performed zero-shot experiments (AUC 78.9). On the RSNA Pneumonia datasets, our CLS model with a linear probe (AUC 90.7) performs slightly worse, but still competitive to results reported by Zhang et al. (2020) and Han et al. (2021). Zhang et al. report CLS combined with linear probing LP (AUC 92.1) and CLS combined with FT (AUC 92.7), whereas Han et al. worked on plain FT (AUC 92.3).
Furthermore, we ran a series of ablations to assess the impact of limited dataset size on the quality of the domain adaptation. We find that CLS can be superior to FT even with only 20,000 image-text pairs (10 % of the MIMIC-CXR dataset). Although, CLS becomes less performant when using fewer than 20,000 training pairs.
Generative Multi-Modal Models open the door to Interpretable and Explainable AI
As shown before, multi-modal models can solve downstream tasks, like classification or retrieval, in a zero-shot manner. Switching to a setting that leverages the generative power of these models opens the door to new and innovative applications. The figure below shows images of chest ray scans for four diagnoses: atelectasis, cardiomegaly, consolidation, and edema. While you can see images generated by our adapted CLIP models in the first row, the second row depicts real radiography images of MIMIC-CXR. Below, we show sentences of the report relevant to the labeling of the diagnosis. Those sentences have been used as prompts for the generative process. To learn more about prompting, check out our prompt engineering guide.
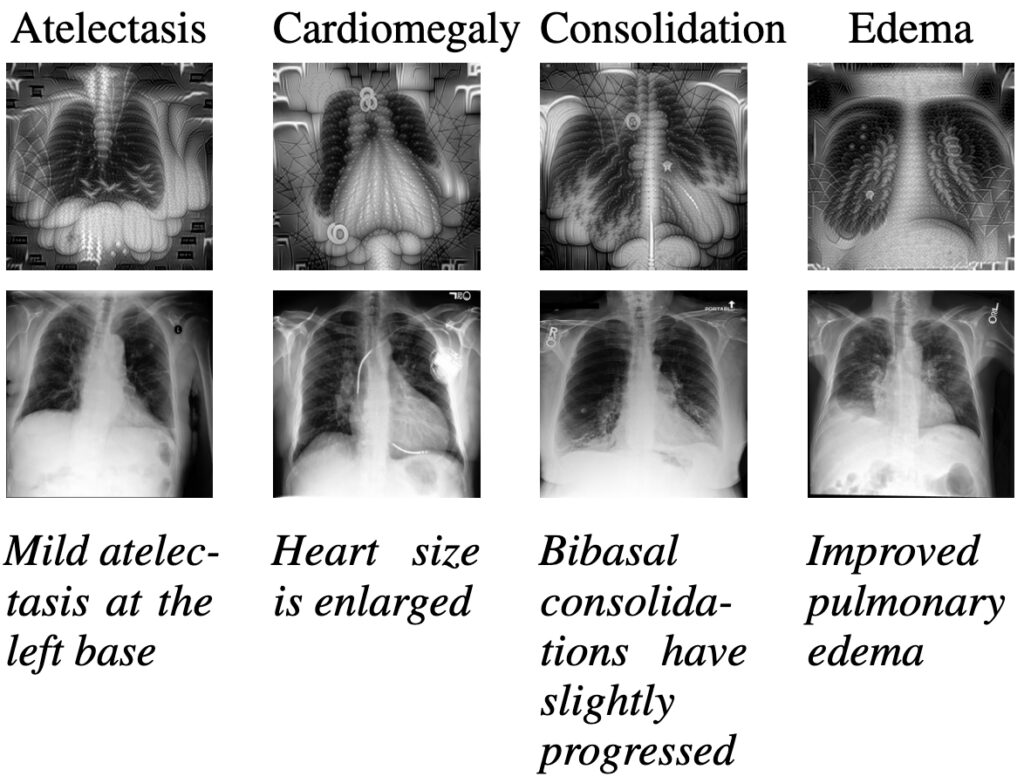
To the inexperienced eye, the generated images share important characteristics with the real X-ray scans. For example, one could argue that lungs and hearts can clearly be seen. When shown to clinicians, such visualizations open the door for qualitative analysis and further empirical studies on interpretable and explainable AI. Potentially, such insights could complement the work of medical experts and are an interesting research endeavour from a medical and AI viewpoint.
Conclusion
We show that Contrastive Language Supervision can be used to adapt CLIP to the domain of chest X-ray scans, as the original features of pre-trained CLIP do not transfer well to this domain. Furthermore, we find that using contrastive language supervision (CLS) with a simple linear probe outperforms fine-tuning (FT) on the MIMIC-CXR dataset and generalizes to other datasets of the same domain. Eventually, we discuss some visualizations generated by our domain adapted models which could be used as a potential building block of future explainable AI systems in medical contexts.
More details can be found in our paper: Language over Labels: Contrastive Language Supervision Exceeds Purely Label-Supervised Classification Performance on Chest X-rays
Further reading
- Domain Adaptation in Natural Language Processing
- Our Prompt Engineering Guide
- Information Extraction from Medical Documents
- Multi-Modal Modeling on vehicle licenses and receipts
- Our E-Health services



