Notice:
This post is older than 5 years – the content might be outdated.
With the sustained success of the Spark data processing platform even data scientists with a strong focus on the Python ecosystem can no longer ignore it. Fortunately, it is easy to get started with PySpark—the official Python API for Spark—due to millions of word count tutorials on the web. In contrast to that, resources on how to deploy and use Python packages like Numpy, Pandas, Scikit-Learn in isolated environments with PySpark are scarce. A nice exception to that is a blog post by Eran Kampf. Being able to install your own Python libraries is especially important if you want to write User-Defined-Functions (UDFs) as explained in the blog post Efficient UD(A)Fs with PySpark.
For most Spark/Hadoop distributions, Cloudera in my case, there are basically two options for managing isolated environments:
- You give all your data scientists SSH access to all your cluster’s nodes and let them do whatever they want like installing virtual environments with virtualenv or conda as detailed in the Cloudera documentation.
- Your sysadmins install Anaconda Parcels using the Cloudera Manager Admin Console to provide the most popular Python packages in a one size fits all fashion for all your data scientists as described in a Cloudera blog post.
Both options have drawbacks which are as severe as they are obvious. Do you really want to let a bunch of data scientists run processes on your cluster and fill up the local hard-drives? The second option is not even a real isolated environment at all since all your applications would use the same libraries and maybe break after an update of a library.
Therefore, we need to empower our data scientists developing a predictive application to manage isolated environments with their dependencies themselves. This was also recognized as a problem and several issues (SPARK-13587 & SPARK-16367) suggest solutions, but none of them have been integrated yet. The most mature solution is actually coffee boat, which is still in beta and not meant for production. Therefore, we want to present here a simple but viable solution for this problem that we have been using in production for more than a year.
So how can we distribute Python modules and whole packages on our executors? Luckily, PySpark provides the functions sc.addFile and sc.addPyFile which allow us to upload files to every node in our cluster, even Python modules and egg files in case of the latter. Unfortunately, there is no way to upload wheel files which are needed for binary Python packages like Numpy, Pandas and so on. As a data scientist you cannot live without those.
At first sight this looks pretty bad but thanks to the simplicity of the wheel format it’s not so bad at all. So here is what we do in a nutshell: For a given PySpark application, we will create an isolated environment on HDFS with the help of wheel files. When submitting our PySpark application, we copy the content of our environment to the driver and executors using sc.addFile. Simple but effective.
Generating the environment
In order to create our aforementioned environment we start by creating a directory that will contain our isolated environment, e.g. venv, on our local Linux machine. Then we will populate this directory with the wheel files of all libraries that our PySpark application uses. Since wheel files contain compiled code they are dependent on the exact Python version and platform.
For us this means we have to make sure that we use the same platform and Python version locally as we are gonna use on the Spark cluster. In my case the cluster runs Ubuntu Trusty Linux with Python 3.4. To replicate this locally it’s best to use a conda environment:
|
1 2 3 |
conda create -n py34 python=3.4 source activate py34 |
Having activated the conda environment, we just use pip download to download all the requirements of our PySpark application as wheel files. In case there is no wheel file available, pip will download a source-based tar.gz file instead but we can easily generate a wheel from it. To do so, we just unpack the archive, change into the directory and type python setup.py bdist_wheel. A wheel file should now reside in the dist subdirectory. At this point one should also be aware that some wheel files come with low-level Linux dependencies that just need to be installed by a sysadmin on every host, e.g. python3-dev and unixodbc-dev.
Now we copy the wheel files of all our PySpark application’s dependencies into the venv directory. After that, we unpack them with unzip since they are just normal zip files with a strange suffix. Finally, we push everything to HDFS, e.g. /my_venvs/venv, using hdfs dfs -put ./venv /my_venvs/venv and make sure that the files are readable by anyone.
Bootstrapping the environment
When our PySpark application runs the first thing we do is calling sc.addFile on every file in /my_venvs/venv. Since this will also set the PYTHONPATH correctly, importing any library which resides in venv will just work. If our Python application itself is also nicely structured as a Python package (maybe using PyScaffold) we can also push it to /my_venvs/venv. This allows us to roll a full-blown PySpark application and nicely separate the boilerplate code that bootstraps our isolated environment from it.
Let’s assume our PySpark application is a Python package called my_pyspark_app. The boilerplate code to bootstrap my_pyspark_app, i.e. to activate the isolated environment on Spark, will be in the module activate_env.py. When we submit our Spark job we will specify this module and specify the environment as an argument, e.g.:
|
1 2 3 4 5 6 7 |
PYSPARK_PYTHON=python3.4 /opt/spark/bin/spark-submit --master yarn --deploy-mode cluster \ --num-executors 4 --driver-memory 12g --executor-memory 4g --executor-cores 1 \ --files /etc/spark/conf/hive-site.xml --queue default --conf spark.yarn.maxAppAttempts=1 \ activate_env.py /my_venvs/venv |
Easy and quite flexible! We are even able to change from one environment to another by just passing another HDFS directory. Here is what activate_env.py looks like, which does the actual heavy lifting with sc.addFile:
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 |
""" Bootstrapping an isolated environment for `my_pyspark_app` on Spark """ import os import sys import logging from pyspark.context import SparkContext from pyspark.sql import SparkSession from pyspark.sql.functions import * _logger = logging.getLogger(__name__) def list_path_names(path): """List files and directories in an HDFS path Args: path (str): HDFS path to directory Returns: [str]: list of file/directory names """ sc = SparkContext.getOrCreate() # low-level access to hdfs driver hadoop = sc._gateway.jvm.org.apache.hadoop path = hadoop.fs.Path(path) config = hadoop.conf.Configuration() status = hadoop.fs.FileSystem.get(config).listStatus(path) return (path_status.getPath().getName() for path_status in status) def distribute_hdfs_files(hdfs_path): """Distributes recursively a given directory in HDFS to Spark Args: hdfs_path (str): path to directory """ sc = SparkContext.getOrCreate() for path_name in list_path_names(hdfs_path): path = os.path.join(hdfs_path, path_name) _logger.info("Distributing {}...".format(path)) sc.addFile(path, recursive=True) def main(args): """Main entry point allowing external calls Args: args ([str]): command line parameter list """ # setup logging for driver logging.basicConfig(level=logging.DEBUG, stream=sys.stdout) _logger = logging.getLogger(__name__) _logger.info("Starting up...") # Create the singleton instance spark = (SparkSession .builder .appName("My PySpark App in its own environment") .enableHiveSupport() .getOrCreate()) # For simplicity we assume that the first argument is the environment on HDFS VENV_DIR = args[0] # make sure we have the latest version available on HDFS distribute_hdfs_files('hdfs://' + VENV_DIR) from my_pyspark_app import main main(args[1:]) def run(): """Entry point for console_scripts """ main(sys.argv[1:]) if __name__ == "__main__": run() |
It is actually easier than it looks. In the main function we initialize the SparkSession for the first time so that later calls to the session builder will use this instance. Thereafter, the passed path argument when doing the spark-submit is extracted. Subsequently, this is passed to distribute_hdfs_files which calls sc.addFile recursively on every file to set up the isolated environment on the driver and executors. After this we are able to import our my_pyspark_app package and call, for instance, its main method. The following figure illustrates the whole concept:
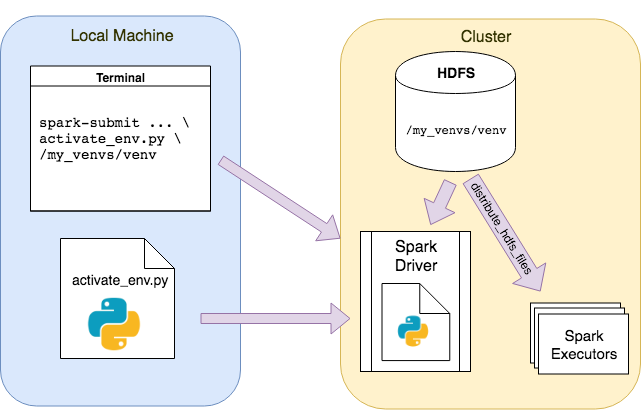
Conclusion
Setting up an isolated environment like this is a bit cumbersome and surely also somewhat hacky. Still, in our use-case it served us quite well and allowed the data scientists to set up their specific environments without access to the cluster’s nodes. Since the explained method also works with Jupyter this is not only useful for production but also for proof-of-concepts. That being said, we still hope that soon there will be an official solution by the Spark project itself.
This article first appeared on Florianwilhelm.info.



