Notice:
This post is older than 5 years – the content might be outdated.
According to the Data Science Survey conducted by JetBrains in 2018, Jupyter/IPython notebooks are the most popular tool in the category IDEs and Editors among data scientists. Whether notebooks can be seen as true IDEs is debatable, however, there is no doubt that they offer several advantages – especially for exploratory data analysis. Incremental execution of code, quick access to visualizations as well as capturing transient computations and the ease of sharing results with others are the major success factors of notebooks.
As for today, there are above 100 compatible kernels, enabling programming in various languages, like Python, Julia, R or even c#. But notebooks offer much more. It is possible to use them for data science on distributed resources, for example by using notebooks in conjunction with a well-known and widely used framework for distributed data processing like Apache Spark. However, traditional notebooks are limited in accessing distributed cluster resources. Furthermore, while working with Spark in combination with Notebooks several challenges must be overcome, especially regarding scalability and stability. Why? Well, it all comes down to the kernel, the unit responsible for interpreting and executing code from the notebook cells.
A traditional notebook container includes the notebook UI and also the kernel itself. Meaning: the notebook and its kernel are running as one unit. To understand the impact this has on working with Spark we first have to take a look at a typical Spark application:
In general, each application runs as an independent group of processes within the cluster and is coordinated by the SparkContext object which exists in the Driver Program. The SparkContext connects to the Cluster Manager (E.g. YARN or Mesos for Hadoop), which then allocates needed resources for the application. Finally, SparkContext provides the Executors with the application code, e.g. as Python code or JAR artefacts and sends tasks to execute.
Now, Spark itself offers two possible deploy modes: Cluster and Client Mode. The main difference between them is where the Spark Driver Program runs. In Client Mode, it is launched directly within the Spark-Submit process, and so it lives within the client, e.g. within a notebook container. In Cluster Mode, the Spark driver is launched outside the client. Speaking of a regular YARN/Hadoop cluster, the Spark driver runs in the application master inside its own YARN container on one of the cluster nodes. After deployment, the client is no longer necessary for further execution. The Spark driver, in contrast to Client Mode, is then subject to the resource manager (YARN) and can be scheduled automatically on nodes with available resources.
After discussing Spark deploy modes, we can now have a look at the combination of both: Spark and Jupyter notebooks. As previously mentioned, in a traditional Jupyter notebook, the kernel exists within the notebook container.

Looking at the image above and remembering how Spark works and what Client and Cluster Mode are, it is obvious that this notebook architecture only allows us to work with Spark in Client Mode. The Spark-Driver and SparkContext are located in the notebook kernel, which is bound to the client process. Unfortunately, the above combination of Spark and notebooks is not flawless. There are several issues that can be improved.
The drawbacks of Spark in Client-Mode within a notebook

Let’s take a look at a common (on-premises) setup consisting of a Hadoop/YARN cluster, as depicted in the illustration above. While Spark-Executors are scheduled across all available cluster nodes by YARN, the Spark driver is not. Instead, it is coupled to the corresponding notebook process (bound to the kernel). When working in a multi-user setup, these heavy notebook processes are utilizing the resources of only one server – usually the cluster edge-node. This dramatically limits the number of users working simultaneously and results in poor scalability.
Stability and user-encapsulation are suffering as well. Since the notebook is not subject to YARN, it is not strictly limited in terms of resources. A common Spark operation of collecting a distributed data-frame back to a notebook (to the Spark driver), e.g. by creating a local (pandas) data-frame, can lead to a quick growth of memory usage. This may exhaust the resources of the machine running the notebook process in no time, possibly freezing the whole machine. In result, all other users‘ processes are negatively affected.
These drawbacks are especially critical when aiming to enable multiple users working on a single platform simultaneously for data science/engineering tasks. A scenario in which one user effectively kills processes/applications of other users is definitely something you want to avoid. But is there a solution to this problem? Yes! The main idea is to separate the notebook kernel from its container with Jupyter Enterprise Gateway.
Remote kernels with Jupyter Enterprise Gateway
Jupyter Enterprise Gateway (JEG) is a web server that enables launching kernels on behalf of remote notebooks. This results in a separation between view and computation since the kernel does not have to run on the same machine (or container) as the notebook like in the traditional setup.

As seen, JEG is used to manage the communication between a notebook and its kernel. In the presented illustration, the notebook does not have multiple kernels running but rather one kernel that is distributed across cluster-nodes. This could be a regular Python kernel that is distributed but also e.g. a kernel containing a Spark application with a Spark driver instance and multiple Spark executors.
The first main advantage of having remote kernels with JEG is improved scalability. Since the kernel does not reside in the notebook, the notebook container requires only minimal resources as it does not perform any computation. In fact, the kernel is now completely schedulable by YARN (or another resource manager), allowing the use of Spark in cluster mode.
With JEG, stability and user encapsulation benefit as well. Because the Spark driver is running as YARN container (in case of a Hadoop cluster) it is strictly constrained in terms of resources as it is managed by YARN. Thinking of the use case mentioned above where a collected Spark data frame could possibly exceed reserved memory, the application will not grow out of bounds but would rather simply fail. While still painful for the affected user, it will not have a negative effect on other users as their Spark drivers might run on another cluster node.
To actually utilize remote kernels in a simplified use case, we need to provide two components: a JEG instance and our actual notebook. Below, we discuss a brief instruction for installing JEG on a Hadoop/YARN cluster to execute PySpark code from a remote notebook. Later we will take a look at a more complex scenario – integrating JEG into a data science platform offering various functionalities, e.g. creating kernels with customized environments within a few clicks from a Web-UI.
Jupyter Enterprise Gateway – Spark in YARN Cluster Mode
We will take a look at a rather basic scenario for remote kernels in which JEG runs on an edge node of a Hadoop cluster and is used to spawn and manage Python Spark kernels. The kernel specification discussed here assumes YARN as a resource manager.
Installing JEG is rather straightforward. It can be done using Conda with the following command executed directly on the edge node:
|
1 |
conda install -c conda-forge jupyter_enterprise_gateway |
If you want to modify JEG’s config, for example to change communication timeouts or specify the amount of allowed kernels for one user, you can use:
|
1 |
jupyter enterprisegateway --generate-config |
In this generated file, you can add various parameters. Click here to visit the official JEG website for more information. After modifying the config-file to suit your needs, it has to be referred while starting JEG.
As kernels are now managed by JEG, we have to provide pre-defined kernels. Per default, JEG discovers available kernels in /usr/local/share/jupyter/kernels/. Make sure this directory has the correct permissions so that JEG is able to access it. While installing and starting JEG is rather easy, the major part of the configuration happens at the kernel level. As already said, we focus on enabling Spark in Cluster Mode.

In the kernel directory, every subdirectory represents one kernel. The file modified the most is the kernel.json. It contains all relevant metadata, like display name (visible from the JupyterLab UI) or a kernel’s programming language. The env section is actually the one that requires most of our attention and an example of this part is presented below:
|
1 2 3 4 5 6 7 8 9 |
"env": { "HADOOP_CONF_DIR": "/usr/bin/hadoop/etc/hadoop", "SPARK_HOME": "/usr/lib/spark", "SPARK_CONF_DIR": "usr/lib/spark/conf", "PYSPARK_PYTHON": "/usr/local/share/jupyter/kernels/py37_folder/py37/bin/python", "PYTHONPATH": "/usr/local/share/jupyter/kernels/py37_folder/py37/bin/python", "SPARK_OPTS": "--master yarn --deploy-mode cluster --name ${KERNEL_ID:-ERROR__NO__KERNEL_ID} --conf spark.yarn.submit.waitAppCompletion=false --conf spark.yarn.appMasterEnv.PYSPARK_PYTHON=py37/py37/bin/python --conf spark.yarn.appMasterEnv.PATH=.py37/py37/bin:$PATH ${KERNEL_EXTRA_SPARK_OPTS} --conf spark.yarn.dist.archives=/usr/local/share/jupyter/kernels/py37_folder/py37.zip#py37 --driver-memory=1024M --num-executors=1 --executor-memory=1024M --executor-cores=1", "LAUNCH_OPTS": "" } |
It contains relevant parameters of a Spark application, e.g. SPARK_HOME or HADOOP_CONF_DIR locations. In SPARK_OPTS we specify the actual deploy mode, resources and the environment (extra packages) we use. Besides these rather basic Spark-related configurations, dependency management is an important aspect. If we want to use e.g. Pandas within our Python-Notebook in a cluster-scenario, we have to make sure that Pandas is installed on all involved cluster nodes. This can be done by creating a custom Conda environment and distributing the archived environment via SPARK_OPTS to all cluster nodes at kernel start. For further information, this blog post is a great read. We install our Conda environment in the kernel directory (conda_environment) and also place the archived environment (conda_environment.zip) here.
To complement our kernel definition, other necessary kernel files like run.sh or the launcher script do not require any changes (at least in the standard scenario) and can be adopted from the provided default kernel, e.g. from here.
After having JEG installed and a prepared kernel available, we can start the JEG server with:
|
1 2 3 |
jupyter enterprisegateway --ip=0.0.0.0 --port_retries=0 --port=8888 --config=' --EnterpriseGatewayApp.yarn_endpoint=http:// |
The YARN endpoint should point to the server, where the YARN master resides. Depending on your cluster this address path may look different.
Once JEG is running on your Hadoop/YARN cluster and is able to discover at least one specified kernel, you are ready to actually connect a notebook started on your local machine (remember the firewall!) with JEG. You simply have to provide the address of JEG (usually the IP & Port of the cluster node on which JEG is running) to your JupyterLab/Notebook at start. You can refer to the official documentation for further details but a simple docker-based command could look like this:
|
1 2 3 4 5 6 7 8 9 10 |
docker run -t --rm -e JUPYTER_GATEWAY_URL='http:// -e JUPYTER_GATEWAY_HTTP_USER=guest -e JUPYTER_GATEWAY_HTTP_PWD=guest-password -e JUPYTER_GATEWAY_VALIDATE_CERT='false' -e LOG_LEVEL=DEBUG -p 8888:8888 -v ${HOME}/notebooks/:/tmp/notebooks -w /tmp/notebooks notebook-docker-image |
Containerized Jupyter Enterprise Gateway
While JEG can simply be started directly on a cluster/edge node, it might be more convenient to have a containerized version including all dependencies and providing more flexibility when thinking about deployments in multiple environments. Of course, you could just use the provided elyra/enterprise-gateway-demo docker image, but if you want it a bit more customized and e.g. use different Spark and/or Hadoop versions (in case you want to use a Spark-related kernel with JEG) than in the provided image, you could also create your own custom docker image.
With JEG running in a container and therefore kernels being started from within the container, we have to include Spark and Hadoop binaries into the container. This can be tricky but the provided Dockerfile brings a basic setup with ubuntu:20.04 as a base image and installation commands for:
- JEG – obviously. It is important to add an entry-point script with a starting command for JEG.
- Hadoop – since JEG needs to access the YARN resource manager that comes with Hadoop. You should install the same or a similar major version as on your cluster.
- Spark – since we want to start Spark applications with our kernels. Again, install the same or a similar major version as on your cluster.
However, simply installing these components will not do it. You actually want to use the physical cluster resources and do not want your Spark application being bound to the resources of the docker host only. The trick is to mount all relevant Hadoop and Spark config-files with volumes into the container at launch. You just have to make sure that you mount the volumes at the right location. E.g. yarn-site.xml from the docker host has to be mounted at its counterpart directory from the Hadoop installed in the container. By mounting all necessary config-files you actually make sure that the (Spark) kernel started inside the container by JEG is able to access the underlying physical cluster resources. Additionally, we also mount the kernel directory enabling us to add new kernels that are immediately recognized by JEG at runtime.
You should also declare variables in your Dockerfile to allow flexible versioning and specify installation paths for Hadoop, Spark or Conda/Python. A build command may then look like this:
|
1 2 3 4 5 6 7 8 9 10 11 12 |
docker build -t --build-arg GATEWAY_VERSION=2.2.0 --build-arg GATEWAY_PORT=8888 --build-arg SPARK_HOME=/usr/lib/spark --build-arg HADOOP_HOME=/usr/bin/hadoop --build-arg PYTHON_HOME=/opt/conda/default/bin/python --build-arg KERNEL_FOLDER=usr/local/share/jupyter/kernels/ --build-arg PYTHON_VERSION=3.7 --build-arg SPARK_VERSION=2.4.6 --build-arg HADOOP_VERSION=2.7 . |
After building (and eventually pushing) the image, you can use e.g. docker-compose to start your container. A YAML-file for Docker-Compose may look similar like the one presented below, however, that depends on the underlying infrastructure you use:
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 |
version: "3.3" services: gateway: image: container_name: jeg-container network_mode: "host" volumes: - /usr/local/share/jupyter/kernels/:/usr/local/share/jupyter/kernels/ - /usr/lib/hadoop/etc/hadoop/yarn-site.xml:/usr/bin/hadoop/etc/hadoop/yarn-site.xml - /usr/lib/hadoop/etc/hadoop/core-site.xml:/usr/bin/hadoop/etc/hadoop/core-site.xml - /hadoop/yarn/:/hadoop/yarn/ - /usr/lib/spark/conf/spark-defaults.conf:/usr/lib/spark/conf/spark-defaults.conf - /usr/lib/spark/conf/spark-env.sh:/usr/lib/spark/conf/spark-env.sh - /usr/lib/spark/jars/:/usr/lib/spark/jars/ - /usr/local/share/google/dataproc/lib/:/usr/local/share/google/dataproc/lib/ - ~/masterarbeit-rafal/jupyter_enterprise_gateway_config.py:/tmp/jupyter_enterprise_gateway_config.py |
You might have noticed that we use host networking for our container. This is related to the way JEG communicates with kernels. At kernel start, JEG maintains a unique response address to which a kernel sends back its status to JEG. This results in multiple open response addresses when running more than one kernel at the same time. This would, among others, require us to expose multiple ports of our container. Ports that we actually do not know in advance. This can be avoided by using host networking. As of JEG version 2.4 single-response addresses for communicating with kernels are not supported, but there is already an open issue on GitHub and we hopefully see this in version 3.0.
This basic scenario allows utilizing remote kernels on your Hadoop/YARN cluster which provides the advantages of Spark in Cluster Mode and the flexibility of programming in a notebook. However, this is rather a single-user solution as there is no management of kernels and it is definitely not an enterprise use case. Ideally, we are able to let multiple data scientists/engineers or developers work with notebooks and utilize shared cluster resources at the same time. To accomplish this, combining JupyterHub with JEG is a good way to go. In this scenario, JupyterHub takes care of spawning single-user notebooks, whereas JEG handles starting kernels and acts as a Gateway. At inovex, we implemented a proof of concept of a holistic data science platform that integrates JEG and offers various functionalities.
Jupyter Enterprise Gateway integrated in a data science platform
The architecture of the proof of concept basically comprises two major parts. An (existing) Hadoop/Yarn cluster for computation loads and a rather small Kubernetes cluster for user interactions. To enable scalability in a multi-user setup we not only distribute the kernels (across the Hadoop/Yarn cluster) but also the actual notebook servers across the Kubernetes cluster nodes!
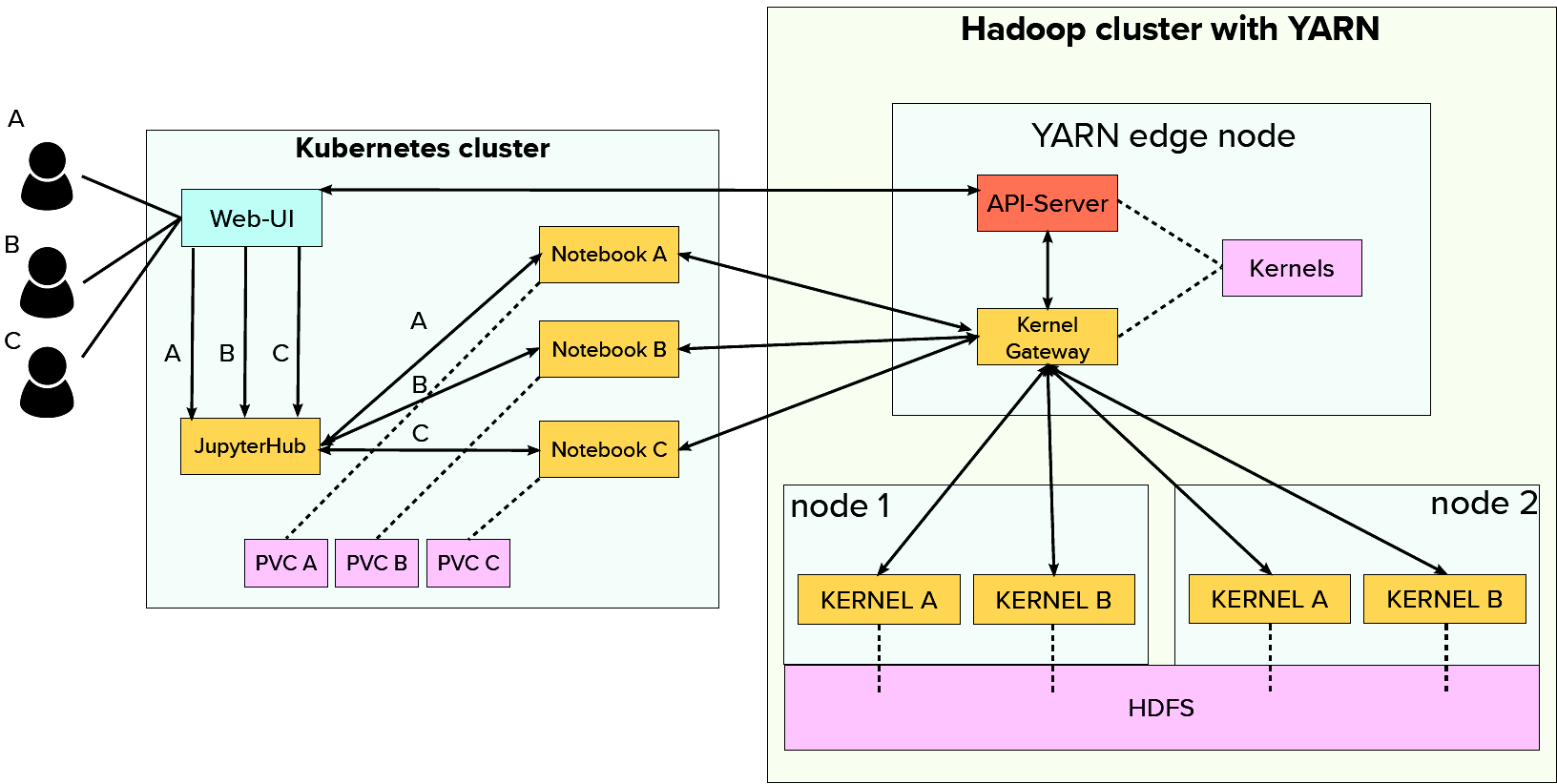
The Web UI running inside Kubernetes acts as a single entry point for the whole platform. After logging in, users are offered various functionalities. They are e.g. able to access JupyterHub and start a JupyterLab session (notebook in the above illustration). JEG (Kernel Gateway) provides the kernels and spawns them on the Hadoop cluster. In this way, Kubernetes is solely responsible for display logic while Hadoop handles the heavy workloads.
Other main functionalities of the Web UI are viewing the cluster state (Hadoop), creating new kernels or setting various resource limits for users from an administrator panel. Since the Web UI is deployed on Kubernetes and JEG as well as kernels residing on the Hadoop cluster, a custom API-Server is needed in case a user wants to e.g. add or modify kernels. It allows the Web UI to communicate with the Hadoop cluster and implements the kernel management functionalities triggered from the Web UI.
The screenshot below presents the core functionalities of the Web-UI. In general, the design is simplistic and focuses on minimizing the technical knowledge required from a user.

The above image is the page a user gets presented immediately after logging in. In the middle, various cluster metrics have been provided. On the left side, you can see all available kernels. They are clickable and lead to the details view, just as depicted below.

Further, the interface for creating new kernels, illustrated below, requires only minimal input from the user. The whole complexity of creating the kernel definition as well as creating & archiving Conda environments is done in the background by the API server.

The interfaces implemented in the Web UI were designed to be possibly error-resistant. E.g. specifying resources beyond limits set by an administrator or exceeding the physical resources leads to an appropriate error message. The same happens by providing the wrong Pip or Conda packages in the case of a Python kernel.
Summary
In this blog post, we have dealt with remote kernels for Jupyter Notebooks and discussed their benefits when working on shared cluster resources. They offer several advantages over a traditional notebook setup in terms of scalability and stability, especially in combination with Spark as a Framework for distributed data processing. Jupyter Enterprise Gateway, a tool from the Jupyter stack, is a good choice to enable remote kernels, whether for a single-user scenario or for enterprise use cases being integrated into a data science platform with other components. Since a kernel specification takes the major part of the configuration and requires quite a bit of technical expertise, we introduced a proof of concept that takes this complexity from the user and offers notebooks with Spark in Cluster Mode out of the box. It utilizes remote kernels and thus offers good scalability, stability as well as user isolation and encapsulation. The platform is also an example of how to integrate various open-source components in order to provide a possibly smooth user experience for data scientists and/or engineers.



